Plasmid
.svg.png)
A plasmid is a small DNA molecule within a cell that is physically separated from a chromosomal DNA and can replicate independently. They are most commonly found in bacteria as small circular, double-stranded DNA molecules; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that may benefit the survival of the organism, for example antibiotic resistance. While the chromosomes are big and contain all the essential genetic information for living under normal conditions, plasmids usually are very small and contain only additional genes that may be useful to the organism under certain situations or particular conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms.
Plasmids are considered replicons, a unit of DNA capable of replicating autonomously within a suitable host. However, plasmids, like viruses, are not generally classified as life.[1] Plasmids can be transmitted from one bacterium to another (even of another species) via three main mechanisms: transformation, transduction, and conjugation. This host-to-host transfer of genetic material is called horizontal gene transfer, and plasmids can be considered part of the mobilome. Unlike viruses (which encase their genetic material in a protective protein coat called a capsid), plasmids are "naked" DNA and do not encode genes necessary to encase the genetic material for transfer to a new host. However, some classes of plasmids encode the conjugative "sex" pilus necessary for their own transfer. The size of the plasmid varies from 1 to over 200 kbp,[2] and the number of identical plasmids in a single cell can range anywhere from one to thousands under some circumstances.
The relationship between microbes and plasmid DNA is neither parasitic nor mutualistic, because each implies the presence of an independent species living in a detrimental or commensal state with the host organism. Rather, plasmids provide a mechanism for horizontal gene transfer within a population of microbes and typically provide a selective advantage under a given environmental state. Plasmids may carry genes that provide resistance to naturally occurring antibiotics in a competitive environmental niche, or the proteins produced may act as toxins under similar circumstances, or allow the organism to utilize particular organic compounds that would be advantageous when nutrients are scarce.[3]
Properties and characteristics
.svg.png)
The American molecular biologist Joshua Lederberg first introduced the term plasmid in 1952 - originally to describe any bacterial genetic material that exists in an extrachromosomal state for at least part of its replication cycle.[4] Later in 1968, it was decided that the term plasmid should be adopted as the term for extrachromosomal genetic element,[5] and to distinguish it from viruses, the definition was narrowed to genetic elements that exist exclusively or predominantly outside of the chromosome and can replicate autonomously.[6]
In order for plasmids to replicate independently within a cell, they must possess a stretch of DNA that can act as an origin of replication. The self-replicating unit, in this case the plasmid, is called a replicon. A typical bacterial replicon may consist of a number of elements, such as the gene for plasmid-specific replication initiation protein (Rep), repeating units called iterons, DnaA boxes, and an adjacent AT-rich region.[7] Smaller plasmids make use of the host replicative enzymes to make copies of themselves, while larger plasmids may carry genes specific for the replication of those plasmids. A few types of plasmids can also insert into the host chromosome, and these integrative plasmids are sometimes referred to as episomes in prokaryotes.[8]
Plasmids almost always carry at least one gene. Many of the genes carried by a plasmid are beneficial for the host cells, for example: enabling the host cell to survive in an environment that would otherwise be lethal or restrictive for growth. Some of these genes encode traits for antibiotic resistance or resistance to heavy metal, while others may produce virulence factors that enable a bacterium to colonize a host and overcome its defences, or have specific metabolic functions that allow the bacterium to utilize a particular nutrient, including the ability to degrade recalcitrant or toxic organic compounds.[6] Plasmids can also provide bacteria with the ability to fix nitrogen. Some plasmids, however, have no observable effect on the phenotype of the host cell or its benefit to the host cells cannot be determined, and these plasmids are called cryptic plasmids.[9]
Naturally occurring plasmids vary greatly in their physical properties. Their size can range from very small mini-plasmids of less than a 1 kilobase pairs (Kbp), to very large megaplasmids of several megabase pairs (Mbp). At the upper end, little can differentiate between a megaplasmid and a minichromosome. Plasmids are generally circular, however examples of linear plasmids are also known. These linear plasmids require specialized mechanisms to replicate their ends.[6]
Plasmids may be present in an individual cell in varying number, ranging from one to several hundreds. The normal number of copies of plasmid that may be found in a single cell is called the copy number, and is determined by how the replication initiation is regulated and the size of the molecule. Larger plasmids tend to have lower copy numbers.[8] Low-copy-number plasmids that exist only as one or a few copies in each bacterium are, upon cell division, in danger of being lost in one of the segregating bacteria. Such single-copy plasmids have systems that attempt to actively distribute a copy to both daughter cells. These systems, which include the parABS system and parMRC system, are often referred to as the partition system or partition function of a plasmid.
Classifications and types


Plasmids may be classified in a number of ways. Plasmids can be broadly classified into conjugative plasmids and non-conjugative plasmids. Conjugative plasmids contain a set of transfer or tra genes which promote sexual conjugation between different cells.[8] In the complex process of conjugation, plasmid may be transferred from one bacterium to another via sex pili encoded by some of the tra genes (see figure).[10] Non-conjugative plasmids are incapable of initiating conjugation, hence they can be transferred only with the assistance of conjugative plasmids. An intermediate class of plasmids are mobilizable, and carry only a subset of the genes required for transfer. They can parasitize a conjugative plasmid, transferring at high frequency only in its presence.
Plasmids can also be classified into incompatibility group. A microbe can harbour different types of plasmids, however, different plasmids can only exist in a single bacterial cell if they are compatible. If two plasmids are not compatible, one or the other will be rapidly lost from the cell. Different plasmids may therefore be assigned to different incompatibility group depending on whether they can coexist together. Incompatible plasmids normally share the same replication or partition mechanisms.[11]
Another way to classify plasmids is by function. There are five main classes:
- Fertility F-plasmids, which contain tra genes. They are capable of conjugation and result in the expression of sex pili.
- Resistance (R) plasmids, which contain genes that provide resistance against antibiotics or poisons. Historically known as R-factors, before the nature of plasmids was understood.
- Col plasmids, which contain genes that code for bacteriocins, proteins that can kill other bacteria.
- Degradative plasmids, which enable the digestion of unusual substances, e.g. toluene and salicylic acid.
- Virulence plasmids, which turn the bacterium into a pathogen.
Plasmids can belong to more than one of these functional groups.
Vectors
Artificially constructed plasmids may be used as vectors in genetic engineering. These plasmids serve as important tools in genetics and biotechnology labs, where they are commonly used to clone and amplify (make many copies of) or express particular genes.[12] A wide variety of plasmids are commercially available for such uses. The gene to be replicated is normally inserted into a plasmid that typically contains a number of features for their use. These include a gene that confers resistance to particular antibiotics (ampicillin is most frequently used for bacterial strains), an origin of replication to allow the bacterial cells to replicate the plasmid DNA, and a suitable site for cloning.

Cloning
Plasmids are the most-commonly used bacterial cloning vectors.[13] These cloning vectors contain a site that allows DNA fragments to be inserted, for example a multiple cloning site or polylinker which has several commonly used restriction sites to which DNA fragments may be ligated. After the gene of interest is inserted, the plasmids are introduced into bacteria by a process called transformation. These plasmids contain a selectable marker, usually an antibiotic resistance gene, which confer on the bacteria an ability to survive and proliferate in a selective growth medium containing the particular antibiotics. The cells after transformation are exposed to the selective media, and only cells containing the plasmid may survive. In this way, the antibiotics act as a filter to select only the bacteria containing the plasmid DNA. The vector may also contain other marker genes or reporter genes to facilitate selection of plasmid with cloned insert. Bacteria containing the plasmid can then be grown in large amounts, harvested, and the plasmid of interest may then be isolated using various methods of plasmid preparation.
A plasmid cloning vector is typically used to clone DNA fragments of up to 15 kbp.[14] To clone longer lengths of DNA, lambda phage with lysogeny genes deleted, cosmids, bacterial artificial chromosomes, or yeast artificial chromosomes are used.
Protein production
Another major use of plasmids is to make large amounts of proteins. In this case, researchers grow bacteria containing a plasmid harboring the gene of interest. Just as the bacterium produces proteins to confer its antibiotic resistance, it can also be induced to produce large amounts of proteins from the inserted gene. This is a cheap and easy way of mass-producing the protein the gene codes for, for example, insulin.
Gene therapy
Plasmid may also be used for gene transfer into human cells as potential treatment in gene therapy so that it may express the protein that is lacking in the cells. Some strategies of gene therapy require the insertion of therapeutic genes at pre-selected chromosomal target sites within the human genome. Plasmid vectors are one of many approaches that could be used for this purpose. Zinc finger nucleases (ZFNs) offer a way to cause a site-specific double-strand break to the DNA genome and cause homologous recombination. Plasmids encoding ZFN could help deliver a therapeutic gene to a specific site so that cell damage, cancer-causing mutations, or an immune response is avoided.[15]
Disease models
Plasmids were historically used to genetically engineer the embryonic stem cells of rats in order to create rat genetic disease models. The limited efficiency of plasmid-based techniques precluded their use in the creation of more accurate human cell models. However, developments in Adeno-associated virus recombination techniques, and Zinc finger nucleases, have enabled the creation of a new generation of isogenic human disease models.
Episomes
The term episome was proposed by François Jacob and Élie Wollman in 1958 to describe extra-chromosomal genetic material that may replicate autonomously or become integrated into the chromosome.[16][17] The use of the term, however, has diverged since it was first coined as plasmid became the preferred word for autonomously replicating extrachromosomal DNA as proposed in the symposium in 1968 – it was suggested by some that the use of the term episome be abandoned, although others continued to use the term with a shift in meaning.[18][19] In prokaryotes, episome is now used by some to refer to plasmid that is capable of integrating into the chromosome. The integrative plasmids may be replicated and stably maintained in a cell through multiple generations, but always at some stage they exist as an independent plasmid molecule.[20]
In eukaryotes, episomes are used to mean non-integrated extrachromosomal closed circular DNA molecule that may be replicated in the nucleus.[21][22] Viruses are the most common examples of this, such as herpesviruses, adenoviruses, and polyomaviruses, but some are plasmids. Other examples include aberrant chromosomal fragments, such as double minute chromosomes, that can arise during artificial gene amplifications or in pathologic processes (e.g., cancer cell transformation). Episomes in eukaryotes behave similarly to plasmids in prokaryotes in that the DNA is stably maintained and replicated with the host cell. Cytoplasmic viral episomes (as in poxvirus infections) can also occur. Some episomes, such as herpesviruses, replicate in a rolling circle mechanism, similar to bacterial phage viruses. Others replicate through a bidirectional replication mechanism (Theta type plasmids). In either case, episomes remain physically separate from host cell chromosomes. Several cancer viruses, including Epstein-Barr virus and Kaposi's sarcoma-associated herpesvirus, are maintained as latent, chromosomally distinct episomes in cancer cells, where the viruses express oncogenes that promote cancer cell proliferation. In cancers, these episomes passively replicate together with host chromosomes when the cell divides. When these viral episomes initiate lytic replication to generate multiple virus particles, they in general activate cellular innate immunity defense mechanisms that kill the host cell.
Plasmid maintenance
Some plasmids or microbial hosts include an addiction system or postsegregational killing system (PSK), such as the hok/sok (host killing/suppressor of killing) system of plasmid R1 in Escherichia coli.[23] This variant produces both a long-lived poison and a short-lived antidote. Several types of plasmid addiction systems (toxin/ antitoxin, metabolism-based, ORT systems) were described in the literature[24] and used in biotechnical (fermentation) or biomedical (vaccine therapy) applications. Daughter cells that retain a copy of the plasmid survive, while a daughter cell that fails to inherit the plasmid dies or suffers a reduced growth-rate because of the lingering poison from the parent cell. Finally, the overall productivity could be enhanced.
In contrast, virtually all biotechnologically used plasmids (such as pUC18, pBR322 and derived vectors) do not contain toxin-antitoxin addiction systems and thus need to be kept under antibiotic pressure to avoid plasmid loss.
Yeast plasmids
Yeast are organisms that naturally harbour plasmids. Notable plasmids are 2 µm plasmids - small circular plasmids often used for genetic engineering of yeast, and linear pGKL plasmids from Kluyveromyces lactis, that are responsible for killer phenotypes.[25]
Other types of plasmids are often related to yeast cloning vectors that include:
- Yeast integrative plasmid (YIp), yeast vectors that rely on integration into the host chromosome for survival and replication, and are usually used when studying the functionality of a solo gene or when the gene is toxic. Also connected with the gene URA3, that codes an enzyme related to the biosynthesis of pyrimidine nucleotides (T, C);
- Yeast Replicative Plasmid (YRp), which transport a sequence of chromosomal DNA that includes an origin of replication. These plasmids are less stable, as they can get lost during the budding.
Plasmid DNA extraction
As alluded to above, plasmids are often used to purify a specific sequence, since they can easily be purified away from the rest of the genome. For their use as vectors, and for molecular cloning, plasmids often need to be isolated.
There are several methods to isolate plasmid DNA from bacteria, the archetypes of which are the miniprep and the maxiprep/bulkprep.[12] The former can be used to quickly find out whether the plasmid is correct in any of several bacterial clones. The yield is a small amount of impure plasmid DNA, which is sufficient for analysis by restriction digest and for some cloning techniques.
In the latter, much larger volumes of bacterial suspension are grown from which a maxi-prep can be performed. In essence, this is a scaled-up miniprep followed by additional purification. This results in relatively large amounts (several hundreds micrograms) of very pure plasmid DNA.
In recent times, many commercial kits have been created to perform plasmid extraction at various scales, purity, and levels of automation. Commercial services can prepare plasmid DNA at quoted prices below $300/mg in milligram quantities and $15/mg in gram quantities (early 2007).
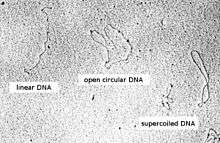
Conformations
Plasmid DNA may appear in one of five conformations, which (for a given size) run at different speeds in a gel during electrophoresis. The conformations are listed below in order of electrophoretic mobility (speed for a given applied voltage) from slowest to fastest:
- Nicked open-circular DNA has one strand cut.
- Relaxed circular DNA is fully intact with both strands uncut, but has been enzymatically relaxed (supercoils removed). This can be modeled by letting a twisted extension cord unwind and relax and then plugging it into itself.
- Linear DNA has free ends, either because both strands have been cut or because the DNA was linear in vivo. This can be modeled with an electrical extension cord that is not plugged into itself.
- Supercoiled (or covalently closed-circular) DNA is fully intact with both strands uncut, and with an integral twist, resulting in a compact form. This can be modeled by twisting an extension cord and then plugging it into itself.
- Supercoiled denatured DNA is like supercoiled DNA, but has unpaired regions that make it slightly less compact; this can result from excessive alkalinity during plasmid preparation.
The rate of migration for small linear fragments is directly proportional to the voltage applied at low voltages. At higher voltages, larger fragments migrate at continuously increasing yet different rates. Thus, the resolution of a gel decreases with increased voltage.
At a specified, low voltage, the migration rate of small linear DNA fragments is a function of their length. Large linear fragments (over 20 kb or so) migrate at a certain fixed rate regardless of length. This is because the molecules 'resperate', with the bulk of the molecule following the leading end through the gel matrix. Restriction digests are frequently used to analyse purified plasmids. These enzymes specifically break the DNA at certain short sequences. The resulting linear fragments form 'bands' after gel electrophoresis. It is possible to purify certain fragments by cutting the bands out of the gel and dissolving the gel to release the DNA fragments.
Because of its tight conformation, supercoiled DNA migrates faster through a gel than linear or open-circular DNA.
Software for bioinformatics and design
The use of plasmids as a technique in molecular biology is supported by bioinformatics software. These programs record the DNA sequence of plasmid vectors, help to predict cut sites of restriction enzymes, and to plan manipulations. Examples of software packages that handle plasmid maps are ApE, Clone Manager, GeneConstructionKit, Geneious, Genome Compiler, LabGenius, Lasergene, MacVector, pDraw32, Serial Cloner, VectorFriends, Vector NTI, and WebDSV. These software help conduct entire experiments in silico before doing wet experiments.[26]
See also
- Bacterial artificial chromosome
- Bacteriophage
- Provirus
- Segrosome
- Transposon
- Triparental mating
- Plasmidome
- DNA recombination
References
- ↑ Sinkovics, J; Harvath J; Horak A. (1998). "The Origin and evolution of viruses (a review)". Acta Microbiologica et Immunologica Hungarica. 45 (3–4): 349–90. PMID 9873943.
- ↑ Thomas, Christopher M; Summers, David (2008). "Bacterial Plasmids". Encyclopedia of Life Sciences. doi:10.1002/9780470015902.a0000468.pub2. ISBN 0470016175.
- ↑ Wolfgang Schumann (2008). "Chapter 1 - Escherichia coli Cloning and Expression Vectors". In Georg Lipps. Plasmids: Current Research and Future Trends. Caister Academic Press. pp. 1–2. ISBN 978-1-904455-35-6.
- ↑ Lederberg J (1952). "Cell genetics and hereditary symbiosis". Physiol. Rev. 32 (4): 403–430. PMID 13003535.
- ↑ Stanley Falkow. "Microbial Genomics: Standing on the Shoulders of Giants". Microbiology Society.
- 1 2 3 Finbarr Hayes (2003). "Chapter 1 - The Function and Organization of Plasmids". In Nicola Casali, Andrew Presto. E. Coli Plasmid Vectors: Methods and Applications. Methods in Molecular Biology. 235. Humana Press. pp. 1–5. ISBN 978-1-58829-151-6.
- ↑ Finbarr Hayes (2003). "Chapter 1 - The Function and Organization of Plasmids". In Nicola Casali, Andrew Preston. E. Coli Plasmid Vectors: Methods and Applications. Methods in Molecular Biology, Vol. 235. Humana Press. pp. 5–6. ISBN 978-1-58829-151-6.
- 1 2 3 T. A. Brown (2010). "Chapter 2 - Vectors for Gene Cloning: Plasmids and Bacteriophages". Gene Cloning and DNA Analysis: An Introduction (6th ed.). Wiley-Blackwell. ISBN 978-1405181730.
- ↑ David Summers (1996). "Chapter 1 - The Function and Organization of Plasmids". The Biology of Plasmids. Wiley-Blackwell; First Edition. pp. 21–22. ISBN 978-0632034369.
- ↑ David P. Clark; Nanette Jean Pazdernik (2012). Molecular Biology (2nd ed.). Academic Cell. p. 795. ISBN 978-0123785947.
- ↑ Margaret C. M. Smith and R. Elizabeth Sockett, eds. (1999). Genetic Methods for Diverse Prokaryotes. Methods in Microbiology, vol. 29. Academic Press. pp. 75–77. ISBN 0-12-652340-1.
- 1 2 Russell, David W.; Sambrook, Joseph (2001). Molecular cloning: a laboratory manual. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory.
- ↑ Uldis N. Streips, Ronald E. Yasbin, eds. (2002). Modern Microbial Genetics (2nd ed.). Wiley-Blackwell. p. 248. ISBN 978-0471386650.
- ↑ Andrew Preston (2003). "Chapter 2 - Choosing a Cloning Vector". In Nicola Casali, Andrew Preston. E. Coli Plasmid Vectors: Methods and Applications. Methods in Molecular Biology, Vol. 235. Humana Press. pp. 19–26. ISBN 978-1-58829-151-6.
- ↑ Kandavelou K, Chandrasegaran S (2008). "Plasmids for Gene Therapy". Plasmids: Current Research and Future Trends. Caister Academic Press. ISBN 978-1-904455-35-6.
- ↑ Morange M (2009). "What history tells us XIX. The notion of the episome" (PDF). Journal of Biosciences. 34 (6): 845–8. doi:10.1007/s12038-009-0098-z. PMID 20093737. (subscription required (help)).
- ↑ Jacob F & Wollman EL (1958), "Les épisomes, elements génétiques ajoutés", Comptes Rendus de l'Académie des Sciences de Paris, 247 (1): 154–156, PMID 13561654
- ↑ Hayes, W (1969). "What are episomes and plasmids?". In Gordon E. W. Wolstenholme; Maeve O'Connor. Bacterial Episomes and Plasmids (CIBA Foundation Symposium). pp. 4–8. ISBN 978-0700014057.
- ↑ Gordon E. W. Wolstenholme; Maeve O'Connor, eds. (1969). Bacterial Episomes and Plasmids (CIBA Foundation Symposium). pp. 244–245. ISBN 978-0700014057.
- ↑ T. A. Brown (2011). Introduction to Genetics: A Molecular Approach. Garland Science. p. 238. ISBN 978-0815365099.
- ↑ Kathleen Van Craenenbroeck, Peter Vanhoenacker and Guy Haegeman (2000). "Episomal vectors for gene expression in mammalian cells". Eur. J. Biochem. 267 (18): 5665–5678. doi:10.1046/j.1432-1327.2000.01645.x. PMID 10971576.
- ↑ Colosimo A1, Goncz KK, Holmes AR, Kunzelmann K, Novelli G, Malone RW, Bennett MJ, Gruenert DC. (2000). "Transfer and expression of foreign genes in mammalian cells" (PDF). Biotechniques. 29 (2): 314–8, 320–2, 324 passim. PMID 10948433. Archived from the original (PDF) on 24 July 2011.
- ↑ Gerdes K, Rasmussen PB, Molin S (1986). "Unique type of plasmid maintenance function: postsegregational killing of plasmid-free cells". Proc. Natl. Acad. Sci. U.S.A. 83 (10): 3116–20. Bibcode:1986PNAS...83.3116G. doi:10.1073/pnas.83.10.3116. PMC 323463
 . PMID 3517851.
. PMID 3517851. - ↑ Kroll J, Klinter S, Schneider C, Voß I, Steinbüchel A (2010). "Plasmid addiction systems: perspectives and applications in biotechnology". Microb. Biotechnol. 3 (6): 634–657. doi:10.1111/j.1751-7915.2010.00170.x. PMC 3815339
 . PMID 21255361.
. PMID 21255361. - ↑ Gunge, N; Murata, K; Sakaguchi, K (July 1982). "Transformation of Saccharomyces cerevisiae with linear DNA killer plasmids from Kluyveromyces lactis". Journal of Bacteriology. 151 (1): 462–4. PMC 220260
 . PMID 7045080.
. PMID 7045080. - ↑ "Vector NTI feedback video". The DNA Lab.
Further reading
- Klein, Donald W.; Prescott, Lansing M.; Harley, John (1999). Microbiology. Boston: WCB/McGraw-Hill.
- Smith, Christopher U. M. Elements of Molecular Neurobiology. Wiley. pp. 101, 111.
- Albert G. Moat; John W. Foster; Michael P. Spector (2002). Microbial Physiology. Wiley-Liss. ISBN 0-471-39483-1.
Episomes
- Piechaczek C, Fetzer C, Baiker A, Bode J, Lipps HJ (1999). "A vector based on the SV40 origin of replication and chromosomal S/MARs replicates episomally in CHO cells". Nucleic Acids Res. 27 (2): 426–428. doi:10.1093/nar/27.2.426. PMC 148196
 . PMID 9862961.
. PMID 9862961. - Bode J; Fetzer CP; Nehlsen K; Scinteie M; Hinrichsen B-H; Baiker A; Piechazcek C; Benham C; Lipps HJ (2001). "The Hitchhiking principle: Optimizing episomal vectors for the use in gene therapy and biotechnology" (PDF). Gene Ther Mol Biol. 6: 33–46.
- Nehlsen K, Broll S, Bode J (2006). "Replicating minicircles: Generation of nonviral episomes for the efficient modification of dividing cells" (PDF). Gene Ther Mol Biol. 10: 233–244.
- Ehrhardt A, Haase R, Schepers A, Deutsch MJ, Lipps HJ, Baiker A (2008). "Episomal vectors for gene therapy". Curr Gene Therapy. 8 (3): 147–161. doi:10.2174/156652308784746440. PMID 18537590.
- Argyros O, Wong SP, Niceta M, Waddington SN, Howe SJ, Coutelle C, Miller AD, Harbottle RP (2008). "Persistent episomal transgene expression in liver following delivery of a scaffold/matrix attachment region containing non-viral vector". Gene Therapy. 15 (24): 1593–1605. doi:10.1038/gt.2008.113. PMID 18633447.
- Wong SP, Argyros O, Coutelle C, Harbottle RP (2009). "Strategies for the episomal modification of cells". Current Opinion in Molecular Therapeutics. 11 (4): 433–441. PMID 19649988.
- Haase R, Argyros O, Wong SP, Harbottle RP, Lipps HJ, Ogris M, Magnusson T, Vizoso Pinto MG, Haas J, Baiker A (2010). "pEPito: a significantly improved non-viral episomal expression vector for mammalian cells" (PDF). BMC Biotechnol. 10: 433–441. doi:10.1186/1472-6750-10-20.
External links
- International Society for Plasmid Biology and other Mobile Genetic Elements
- History of Plasmids with timeline